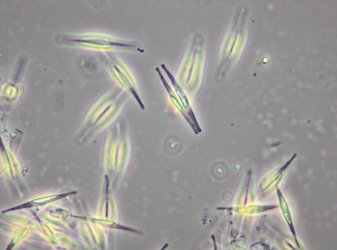
Phaeodactylum tricornutum is a microscopic, single-celled algae with outsized potential.
It is a leading contender to improve sustainable production of biodiesel and other products using seawater and carbon dioxide as raw materials. It captures and stores energy from light, grows quickly and contains a high proportion of lipids, which provide essential oils to much of the marine food chain.
Yet how these diatoms do what they do is not well understood – and that’s where Jamey Young, associate professor of chemical and biomolecular engineering, and his lab come in. He’s part of a team led by the J. Craig Venter Institute (JCVI) that received a five-year, $10.7 million grant from the U.S. Department of Energy to optimize metabolic networks in photosynthetic microalgae.

Engineered microalgae could play a huge role in creating sustainable energy. Based on their growth potential and photosynthetic efficiency, oil-rich microalgae could produce about 200 barrels of biofuel per hectare of land, according to JCVI. That’s 100 times the amount of biodiesel currently produced from soybeans in the United States.
The oil content of Phaeodactylum tricornutum makes it an attractive option; it also is one of only four diatom species for which a complete genome sequence has been generated.
“Diatom photosynthesis is estimated to account for between 25 percent and 40 percent of the nearly 50 billion tons of organic carbon fixed annually in the sea,” said Young , who is also a Vanderbilt Chancellor’s Faculty Fellow, “yet their metabolic pathways have not been rigorously characterized at the systems level.”
“The true potential for light-driven metabolism remains poorly understood,” he said.
The new research effort is expected to develop technologies needed to achieve sustainable production of biochemicals from photosynthetic microbes within the next 10 to 15 years, Young said.
The goal is to leverage significant recent advances in diatom genome engineering and metabolic modeling to dramatically improve the ability to understand and redirect metabolism in these microbes.
In prior DOE-funded research, the Young lab developed a suite of experimental approaches and software packages that enable 13C flux analysis of photosynthetic metabolism in bacteria and plants. This project will leverage those tools to perform 13C flux analysis of the model diatom Phaedoctylum tricornutum. The resulting flux maps will be integrated with genome-scale metabolic models to identify bottlenecks that limit cell growth and/or lipid production, which can be overcome through genome engineering.
This project involves a collaboration between investigators at the JCVI, a non-profit genomic research organization based in La Jolla, CA; Colorado State University; UC-San Diego; and Vanderbilt. The grant is from DOE’s Office of Science, Biological and Environmental Research (BER) Genomic Science Program.
The research is funded through DE-SC0018344: Design, Synthesis, and Validation: Genome Scale Optimization of Energy Flux through Compartmentalized Metabolic Networks in a Model Photosynthetic Eukaryotic Microbe. This award was made in response to solicitation DE-FOA-0001650, Biosystems Design to Enable Next-Generation Biofuels and Bioproducts.
Media Inquiries:
Pamela Coyle, (615) 343-5495
Pam.Coyle@Vanderbilt.edu
Twitter @VUEngineering